Overview
Research Focus
Select Publications
Facilities
Projects
Research Staff
Overview
Computational Engineering is a relatively new discipline that deals with the development and application of computational models and simulations, often coupled with high-performance computing, to solve complex physical problems arising in engineering analysis and design. The research focus of this laboratory is on the use of heterogeneous parallel processing to speed up the bioinformatics applications such as cancer genomics pipelines, data science and analytics, modeling and simulation in medical applications (especially drug discovery and spiking neural networks), and simulation of molecular dynamics for studies in computational sciences. A heterogeneous cluster computing facility has been created to meet the huge computational needs of all these research activities. Cyber security and development of (i) Pulse analyzer device for use in traditional medicine and (ii) Bioinformatics toolbox are the other ongoing research activities.
Research Focus
Cancer Genomics
Genomics is the study of all the genes in the genome and their interactions with the environment. Genomic analysis concerns with bio-molecules such as DNA sequences and RNA sequences in (i) identifying them, (ii) comparing their features and (iii) measuring their structural variation, gene expression, or regulatory and functional element annotation. Next Generation Sequencing (NGS) technologies help in the characterization of point mutations of a wide range of cancers and their structural alterations. The complete genome sequences of numerous cancer types are becoming increasingly available, which provide a comprehensive view of cancer development and growth. Cancer genome studies enable understanding the gene abnormalities that are the root cause of many types of cancer. This improved understanding of cancer biology helps in developing new ways of diagnosis and treatment.
Cancer accounts for approximately 13% of all deaths worldwide and contributes significantly to global socio-economic burden to the society. Evidences have supported the theory that cancer is a disease of genome and aberrant genomic alterations are the hallmark of cancer cells. Current research in cancer genomics is focused mainly on understanding intratumor heterogeneity which plays a major role in tumor metastasis and drug resistance to therapeutics. In order to better understand the tumor genome heterogeneity in tumor niche, a global approach of sequencing the whole cancer genome was undertaken. Since then, several sequencing studies have identified a strong correlation between genomic alterations with tumor progression and resistance to conventional therapies. However, effective treatment of cancer relay on early detection, precise diagnosis and implementation of specific therapeutic strategies, which give rise to a concept of personalized medicine.
Although, whole genomic sequences provide important insights on clonal evolution of mutations, the lack of the proper tools and pipelines to analyze and interpret the framework of data has limited the use of sequencing technology for diagnosis and therapeutics. Independent studies have identified several driver mutations with specific cancer type but analyzing how each of these mutations work individually or in a cohort towards tumor progression is limiting. Recent advancement in computational biology and mathematical modeling has shown great potential in analyzing the huge sequencing data and in understanding the origin and progression of cancer without much of the wet lab experiments and ethical issues.

Genome Analysis Pipeline
A computational model for identifying multiple cancer specific biomarkers from the whole genome sequence data of cancer patient and analyzing cancer prognosis using predictive machine learning models is proposed. The source of the whole genome sequencing data of cancer cells will be obtained from European Genome-Phenome Archive (EGA), The Cancer Genomics Hub (CGHub) and other relevant databases. The source for the potential biomarkers known for each cancer type will be obtained from 1000 genomes browser and COSMIC databases and the reference sequences of these biomarkers will be obtained from National Center for Biotechnology Information (NCBI). The pipeline developed will be a cluster of four main groups based on their application such as assembly, alignment, variation discovery and annotation. For predictive modeling, machine learning approaches will be used, that have unique characteristics for prediction and classification such as capacity control of the decision function, the use of the kernel functions, and the scarcity of the solution. The three-way data split using this model will be applied for training, validation and testing. The proposed pipeline and a schematic representation of the frontend of the pipeline are shown below.
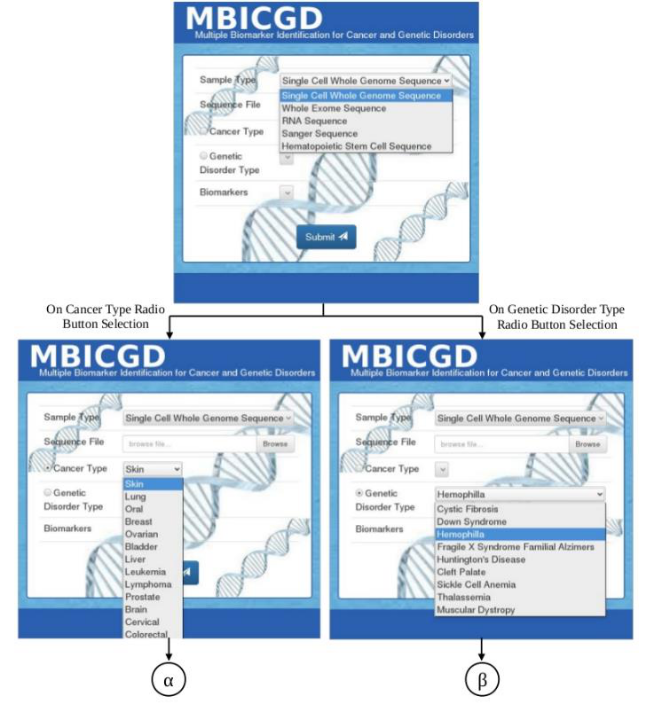

Frontend of the MBICGD Pipeline

Frontend of the MBICGD Pipeline (contd.)
Research Objectives
- To recognize and use the new opportunities offered by NGS technologies in the field of cancer genomics.
- To overcome the challenges raised by the massive amount of sequence data that has been and will be generated.
- To create a dedicated bioinformatics facility for the genome analysis dedicated to cancer studies.
- To empower the biomedical teams working in the field of cancer to best exploit the huge data generated by the ongoing genomic projects.
Research Outcome
- High-throughput tools for DNA and RNA characterization that facilitate comprehensive analyses of cancer genomes including all somatic alterations.
- Cancer genome methodology is to understand the relationship between the genetic mutations and the clinical response to assist in cancer therapy.
- New methodologies for cancer diagnosis and prognosis.
- Comprehensive catalogs of the genomic changes in respect of specific cancer types.
- Technologies for complete molecular profiling of tumors enabling genome-informed personalized cancer medicine.
- New techniques for validation of genomic data to (i) distinguish mutations responsible for disease pathogenesis resulting from genomic instability (ii) define genes responsible for cancer initiation, progression, and maintenance, and (iii) identify the effective ways to therapeutically exploit this information.
Beneficiaries
- Cancer patients
- Medical practitioners
- Genome scientists
- Research scholars
Data Science and Analytics
In most organizations, various types of data are collected in such large volumes that these organizations can no longer process, properly store, or transmit these data over regular communication lines in a timely manner.
As analysts collect huge volume of data and use those data in complex analytical workloads, many organizations, compute and storage silos are proliferating across research groups. In order to overcome the technical challenges in the new era, there is a demand for an architecture which can make it possible for healthcare and life science organizations to easily scale compute and storage resources as demand grows, and to support the wide range of development frameworks and applications required for industry innovation all without unnecessary re-investments in technology and the storage systems must also be designed such that the same storage media can support different access methods in a reliable, yet flexible manner. To address these requirements, a reference architecture has been proposed and will be built based on the best practices of heterogeneous parallel computing technology (as shown in Figure 3).
MapReduce Hadoop Framework is used in large enterprise applications in all domains including healthcare, public sector administration, global personal location services, online retails, manufacturing, social websites, etc. Hadoop requires huge computational resources since it must process ever growing big data. With the advent of heterogeneous parallel programming, many applications are accelerated to achieve high performance by making extensive use of both CPUs and GPUs.
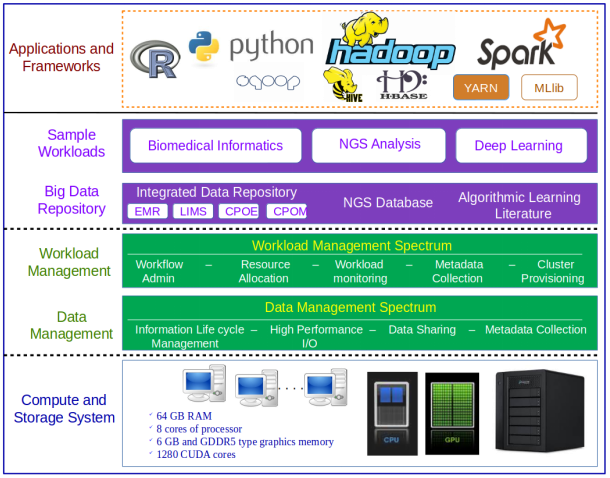
Big Data Analytics Architecture for Healthcare
Research Objectives
- To implement Hadoop framework to optimally utilize CPUs and GPUs in a CPU-GPU distributed cluster for data science and analytics applications.
- To fine tune the distributed cluster for use in data science and analytics projects.
Research Outcome
- Hadoop framework implemented on a heterogeneous parallel processing platform for use in data science and analytics projects.
Beneficiaries
- Large scale enterprises utilizing the big data analytics for their business intelligence.
- Data scientists.
- Researchers
Molecular Level Protein-Ligand Interactions
A ligand is a molecule or an ion which has some affinity towards protein and gets bound to form a complex substrate. Protein-Ligand interactions involve observation of how a ligand acts on a complex protein structure and gets bound for a specific purpose. The process of binding a ligand to a protein is called docking. In this process, the spherical ligands get attached to the twisted helix structured protein. Rendering large and complex structure of molecules and the frame rate required for complex interactions require vast computational resources. Use of heterogeneous multi-core CPUs and many-core GPUs in a cluster promises to mitigate this computational resource crunch.
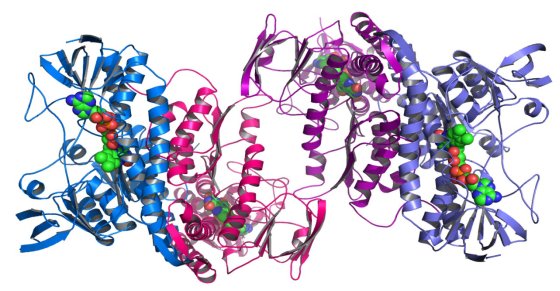
Protein-Ligand Interactions
Research Objectives
- To develop methods to find out the proper orientation of ligand towards protein molecule based on the bonding affinity.
- To use this to dock a ligand to protein and to rescore the docking process using accurate methods to overcome previous dock scoring functions.
- To develop a virtual screening pipeline which gives high throughput and scans virtual compound databases using massively parallel pipeline.
- To find out the specific orientation to match with large sets of complex compounds to improve drug discovery.
- To arrive at anappropriate score function and apply to rescore the affinity
- To create automatic receptor and binder function to identify the binding site.
Research Outcome
- Virtual screening pipeline developed with effective rescoring using massively parallel cores enabling the discovery of better lead compounds.
- System and processes for high speed screening of compounds using massively heterogeneous parallel pipeline to achieve better drug discovery.
Beneficiaries
- Pharmaceutical companies.
- Whole ecosystem of healthcare since effective drugs can be discovered fast.
- Systems biology research community.
Spiking Neural Networks Simulator
Spiking neural networks model improves the level of practicality in the neural simulation. Additionally, the state of neuronal and synaptic timing is integrated into Spatial Statistical Networks (SSNs) working model. This avoids the neurons reaching the circulation cycle but fire when they are membrane prospective. When the neuron fires, a signal is generated which reaches the other neurons and as a result it either increases or decreases the capabilities in relation to the signal. Due to huge computational resource requirements, parallel CPU-GPU co-processing is used in spiking neural simulators that integrate event and time computational methods. Time-driven simulation using CPU architectures works well for small-scale spiking neural simulation but cannot scale to simulate large-scale spiking neural networks. However, event-driven simulation on CPUs and time-driven simulation on GPUs perform better than CPU based time-driven simulation in specific cases. A combined event and time-driven simulator using an amalgamated CPU–GPU platform is being developed to speed up spiking neural networks simulation. This neural simulator will support simulation of different models of bio-inspired spiking neural networks facilitating both event-driven and time-driven schemes (time-driven schemes on CPU and GPU and event-driven schemes on CPU) and will work on different combinations of processing cores in the same simulation.

Spiking Neural Network
Research Objectives
- To integrate different simulation techniques on a single simulation platform supporting time and event driven techniques.
- To conduct performance characterization study using simulation.
- To improve propagation and queue management times of spikes.
- To optimize synchronization and transfer times when using heterogeneous multicore CPU and many core GPU platform.
- To develop methods to switch simulation activities from CPU to GPU by exploiting the computing power of GPU.
- To simulate different models of biologically inspired spiking neural networks.
Research Outcome
- Spiking neural network simulation platform supporting time and event driven techniques.
- Biologically inspired versatile tool for speeding up the simulation of spiking neural networks.
Beneficiaries
- Researchers
Wrist Pulse Analyzer
Wrist pulse analysis is a simple non-invasive technique used in ancient Indian medicine Ayurveda and Traditional Chinese Medicine (TCM) for health diagnosis. Most of the modern pathological diagnostics tools give reports in terms of content and chemical changes in the patient’s body. Very few devices give reports based on body constitution of the patient. Wrist pulse diagnosis is one of the ancient methods of diagnosing human body constitution. Computerized pulse diagnosis uses sensors to acquire the wrist pulse signals and machine learning techniques to analyze patient’s health on the basis of the acquired pulse signals. Wrist pulse indicates the significantly varied blood flow of an organ in abnormal health conditions. Patient health conditions manifest themselves in the reflected waves of the wrist pulse signals in different ways. With careful interpretation of the wrist pulse signals, one can perform an initial diagnosis to a great accuracy.
Research Objectives
- To design an efficient and accurate pulse acquiring device without losing vital radial pulse information of the patient.
- To acquire the radial arterial acoustic vibrations through non-invasive pulse detector.
- To develop an indigenous application to analyze, characterize and classify the human radial pulse obtained through the detector.
- To provide low cost pulse diagnosis system for alternative and integrated medical practitioners.
- To provide a complete analysis of the overall constitution of a person in accordance with the TCM and Ayurveda principles through charts and bar graphs.

Wrist Pulse Analysis
Research Outcome
- A wrist pulse detector which takes the acoustic vibrations of the radial arterial pulse and an intelligent machine which analyses the pulse variations and provides vital five elements energy level information which aids in the correct diagnosis of the ailments.
- Low cost pulse diagnosis system for alternative and integrated medical practitioners.
- A system for the complete analysis of the overall constitution of a person in accordance with the TCM and Ayurveda principles through charts and bar graphs.
Beneficiaries
- Alternative medicine practitioners (AYUSH) for getting the complete body constitution of the patient.
- Patients in terms of getting better and affordable alternative therapies.
- Researchers to pursue their research on pulse diagnosis and alternate therapies.
Whole-Cell Modeling and Simulation
Understanding and engineering of biological systems requires comprehensive models of cellular physiology with 100% predictability. These whole-cell models guide experiments in molecular biologyand enable simulation and computer-aided design in synthetic biology. They help personalized medical treatment. Constructing comprehensive whole cell models with sufficient detail and validating them is a massive work. Modeling larger cells and more complex physiology involve (i) model building and integration, (ii) model validation, (iii) data curation, (iv) experimental interrogation, (v) analysis and visualization, (vi) accelerated computation, (vii) community development and (viii) collaboration. A broader multidisciplinary research community is needed to innovate in all these areas. As a first step, a study on the issues and challenges is initiated and in progress.
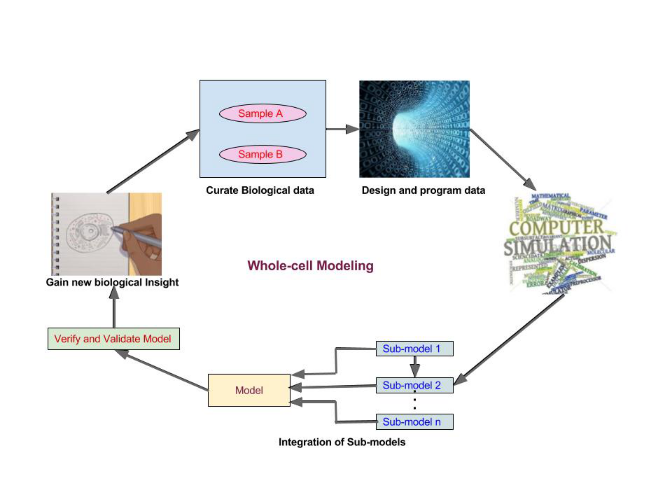
Whole-Cell Modeling Process
Cyber Security
The application of chaos theory in the emerging field of chaotic cryptography has been motivated by the chaotic properties such as ergodicity and sensitive dependence on initial conditions and system parameters, in addition to complex dynamics and deterministic behaviors. A tiny change in the initial values can lead to completely different results. The NIST standard is commonly applied to test the randomness for pseudo-random bit sequence; results demonstrate that a good pseudo-random sequence can be generated by chaos to meet the requirement of image encryption. It gives a novel confusion and diffusion method for image encryption. In the scheme developed for image encryption, the algorithm first shuffles the position of pixel values and then changes the gray values to make the complex relationship between original plain image and encrypted image. Image scrambling and diffusing, both operations are performed by logistic map. Different experiments have been conducted and the results show that the scheme is resistant to different cryptanalytic attacks and provides adequate security.
The primary objective of this research is to enable secure transmission of digital images over unsecured interconnected networks since the communication of digital information and images over network occur more and more frequently nowadays. The specific objectives are:
- To develop the unconventional cryptosystem for image encryption which can provide better security to images as compared traditional cryptographic techniques such as DES, IDEA and RSA.
- To use characteristics of digital chaotic system such as non-periodicity, like-random behavior, sensitivity to build image encryption scheme.
- To provide secure communication as the internetwork and personal communications systems are accessible worldwide.
- To design a new chaotic cryptosystem for obtaining high level security against known cryptanalytic attacks.
Chaotic Map
The simplest chaotic map is logistic map. The Logistic map presents a polynomial map of degree two. It is equivalent to the recurrence relation. It is a simple and one-dimensional discrete-time non-linear system. This widely used function exhibits quadratic non-linearity and is defined by following equation:
Where is a control parameter and it lies between 0 and 4. The chaotic sequence is represented by and it lies between 0 and 1. The logistic map becomes chaotic when the control parameter lies between 3.57 and 4. The bifurcation figure of one-dimensional logistic map has been shown in Figure 1(a) and (b). The diagram presents periodic windows in fixed intervals. The existence of periodic windows must be avoided; otherwise the cipher text would not show random-like behavior and resulting in an inefficient encryption procedure.
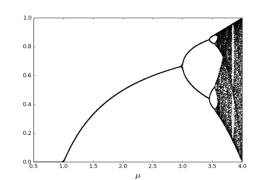
Bifurcation diagram for µ<1
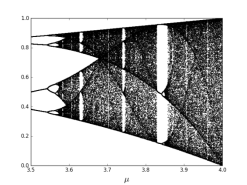
Bifurcation diagram for µ>3.5
Digital technology is growing rapidly, causing a wide spread of distribution of digital documents and images over the Internet. The security of digital documents, images and other multimedia data has thus become extremely important for common people and the government. A secured transmission of digital images is an important issue in information security field. Cryptographic encryption techniques provide an effective security to data by converting it into un-understandable form to attackers. Conventional cryptographic encryption methods are not suitable for image encryption. In recent years chaotic theory has attracted research community for image encryption. A simple and secured scheme for image encryption using one-dimensional logistic maps has been studied. This image encryption scheme first shuffles the position of pixel values and then changes the gray values leading to a complex relationship between the original plain image and the encrypted image. Two operations: image scrambling and diffusing are performed by logistic maps. Various experiments test the robustness and the security aspect of the algorithm and it is found that such schemes are resistant to different cryptanalytic attacks and provides adequate security.
With the rapid growth of multimedia production systems, electronic publishing and widespread dissemination of digital multimedia data over the Internet, protection of digital information against illegal copying and distribution has become extremely important. Encryption on image has its own requirements due to the intrinsic characteristics of images such as bulk data capacity and high redundancy. Traditional symmetric encryption algorithms such as DES, IDEA, Blowfish and RSA are generally not suitable for image encryption due to their slow speed in real-time processing and some other issues such as in handling various data formats.
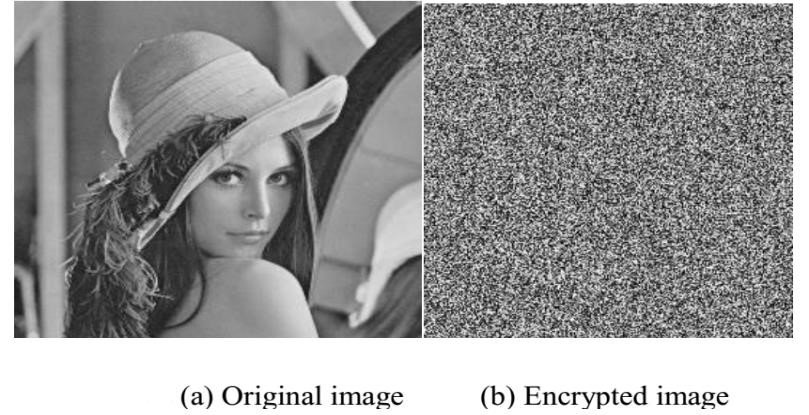
The gray-scale Lena image 256×256 of size is considered for the sake of testing the algorithm. Figure 2(a) and Figure 2(b) shows the original Lena image and corresponding cipher image respectively. Experiments result shows that effect of encryption process is good, as the logistic map generates the random chaotic sequences.

Select Publications
- An Efficient Component Based Filter for Random Valued Impulse Noise Removal. Manohar Koli, Balaji S. International Journal of Applied Engineering Research (2016)11(3):1908-1915.
- Energy Dissipation Model for 4G and WLAN Networks in Smart Phones. Shalini Prasad, Balaji S. International Journal of Advanced Computer Science and Applications (2016)7(7):62-68.
- Advances in Data Mining for Internet of Things. Abhishek K., Balaji S. International Journal of Control Theory and Applications (2017) 10(9):57-66.
- Modeling and Simulation of Cell Biological Systems on Heterogeneous Parallel Computing Platforms: A Review. Aditya Pai H., Sreenivasa N., Balaji S.International Journal of Control Theory and Applications (2017) 10(9):79-89.
- Multi- frame Twin Channel Descriptor for Person Re-identification in real time surveillance videos. Sathish P.K., Balaji S. International Journal of Multimedia Information Retrieval Accepted
- Merkle-Damgård Construction Method and Alternatives: A Review. Harshvardhan Tiwari. Journal of Information and Organizational Sciences- Accepted.
- Recent Trends in Application of Neural Networks to Speech Recognition. Gneswari G., Vijaya Raghava S.R., Thuskar A.K., Balaji S. International Journal of Recent and Innovation Trends in Computing and Communication (2016) 3(1):18-25.
Facilities
The Computational Engineering lab is equipped with a High-Performance Computing Cluster with 11 high-end workstations interconnected using a high-speed backbone interconnect. Each of the workstation possesses an 8-cores CPU with 16-threads capability, 64GB RAM, 1286-cores GPU card, 1 TB of storage and other peripherals. This provides 88 CPU cores, about 700 GB of RAM, about 14,000 GPU cores to the users in the cluster environment that can be used to speed up the compute bound applications in science, engineering and technology. The total processing power of the cluster is about 50 TFLOPS. If required, the cluster memory can be upgraded to 1.4TB without any changes to the hardware. Heterogeneous parallel programming is used to effectively utilize these computing resources.
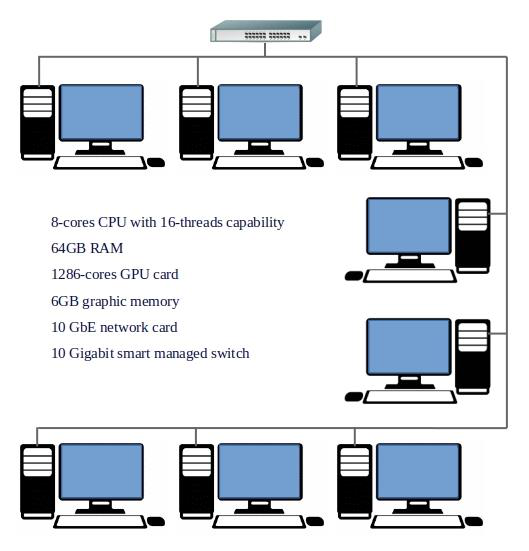
High Performance Computing Cluster
Projects
- Development of Computational Model for Identification of Multiple Biomarkers in Whole Cancer Genome and Predictive Model for Cancer Prognosis- SSPS.
- Development of Mobile Based Objective Type Test Delivery and Analysis Platform- SSPS and S-VYASA University.
- PanchNidaan – A Software Tool for Traditional Patient Diagnosis Using Tongue, Lips, Eyes, Nails and Pulse- SSPS.
- Big Data Analytics Platform for Healthcare- SSPS
Research Staff


Balaji received his Ph.D. in Computer Science and Engineering from Indian Institute of Science, Bengaluru in 1993. The title of his thesis S-NETS: A Tool for the Performance Evaluation of Hard Real-Time Scheduling Algorithms, is on hard real-time systems. His registration for M.Sc.(Engg) in the area of Fault-Tolerant Computing in the same department was upgraded in 1989 to Ph.D. due to the outstanding progress made at graduate level. He holds Masters Degree in Applied Mathematics from Anna University (1981) and a B.Sc. in Mathematics, Physics and Statistics from Madras University (1979).
Balaji is a technologist turned academician with a blend of academic and industrial experience and expertise in successfully executing and managing R&D intensive projects. He has considerable experience in developing mission and safety critical applications and cutting-edge systems in embedded and mobile computing. He also brings vast experience at senior management levels during his stint in ISRO Satellite Centre, country’s premier space research organization. His forte is in ensuring that all the functional units within an organization are aligned to deliver best value to the organization. His strong technical foundation and his experience at senior management levels help him move seamlessly between finance, administration, project management and engineering.
As a technologist, Balaji has several accomplishments. He has defined and established software processes for developing mission and safety-critical space applications. He has developed a number of simulation packages for the design and design validation of attitude and orbit control systems of satellites. He has introduced numerous innovations for the design validation of spacecraft onboard sub-systems and developed advanced test systems to validate the satellite control systems. He has setup the facility for the software development, using high-level programming languages ADA, of processor-based satellite control systems, institutionalized processes and methodologies for development of such systems and has trained/mentored staff to deliver high integrity systems that meets critical spacecraft mission requirements. He has developed several in-house tools for software engineering and project management.
He has served at AMC Engineering College, City Engineering College in various capacities, such as Principal/Vice-principal. He then worked at Centre for Emerging Technologies, Jain University where he initiated research in the areas of bioinformatics, genomics and molecular modelling and simulation.
Balaji has been a member of several professional bodies, such as, ISTE, Astronautical Society of India, Association for Computing Machinery, Institution of Electrical and Electronic Engineers, Institute of Smart Structures and Systems and a fellow of IETE. Balaji’s current research interests include, Heterogeneous parallel processing, Data science and analytics, Bioinformatics, Genomics, Computational sciences, Modeling and simulation, Video data mining and analytics, Image processing, Embedded and real-time computing, Affordable medical devices, Energy harvesting and energy management, Mobile computing.
During his tenure at ISRO Satellite Centre, Balaji was
(i) a member/convener of several software review committees,
(ii) a member/convener of several software verification and validation committees,
(iii) a member of Seminar Committee,
(iv) a member of Library Committee,
(v) Executive Secretary, Editorial Board, Journal of Spacecraft Technology and
(vi) Executive Secretary, Association for Autonomous and Fault-Tolerant Systems. He was the Organizing Secretary of the Information Technology Track of the Indian Science Congress, 2002.
Balaji has delivered invited talks on the recent trends in computer science and engineering at various national conferences and workshops. He was a member of the Technical Committee of a few national conferences. He has been a member of Board of Studies in Computer Science and Engineering in a couple of engineering colleges. He served as the internal representative of all Research Review Committees – Computer Science and Engineering, Jain University during Mar 2014 – Jul 2016. He has been a member of the Comprehensive Examination Committees of VTU and has been a thesis examiner at Anna University. He is an Authorized Research Supervisor in Computer Science and Engineering at Visveswaraya Technological University, Jain University and Tumkur University. He has published over thirty research articles in refereed national and international journals. He has authored chapters in two books published by Springer and CRC Press. He is currently working as a Professor at the Centre.
Email Id: balaji.s@ciirc.jyothyit.ac.in

Harshvardhan Tiwari received his Ph.D degree in Computer Science and Engineering from JIIT University, Noida, Uttar-Pradesh, India. He completed his post-graduate study (MTech) and graduate study (BE) both in Computer Science and Engineering from RGTU, Bhopal, Madhya-Pradesh, India in 2009 and 2005 respectively. His research interests are computer network, algorithms, DBMS, network security and information security. He has more than 4 years of teaching and research experience. He has several publications to his credit that are SCOPUS and DBLP indexed. During his academic career, he pursued and experienced teaching subjects like Distributed Operating System, Computer Architecture and Organization, Database Management System, Cryptography and Information Security, Computer Networks, and Web Technologies. Apart from teaching, he is an active researcher and currently advisor of two research scholars. He is also a member of technical program committee of referred Journals and Conference Proceedings. He is actively involved in curriculum planning, syllabus design at an engineering graduate level. He has also served as an external examiner and adjunct instructor at other Universities. He has participated in different faculty development programs. He is a member of many professional scientific bodies such as IAENG, ICST.
Email Id: harshvardhan.t@ciirc.jyothyit.ac.in

Nayana G Bhat received her Bachelor’s degree in Electrical & Electronics Engineering from Visvesvaraya Technological University in 2004 and M.Tech in Computer Science and Engineering from Kuvempu University in 2006. She has more than eight years of teaching experience in various engineering colleges. She has taught many computer science UG subjects and has guided under graduate projects. She has participated many National conferences, faculty development programs, seminars and workshops. She is a member of CSI, ISTE. Her research interests include parallel computing, image processing, data mining, and neural networks. Nayana is currently working as an Assistant Professor.
Email Id: nayana.gb@ciirc.jyothyit.ac.in
Rashmi K S holds a Bachelor’s degree (2009) and a Master’s degree (2014) in Computer Science and Engineering from Visvesvaraya Technological University. She worked as Assistant Professor in Jain University since 2014 where she taught undergraduate and post-graduate students. Her research interests include bioinformatics, big data analytics, Internet of Things, molecular modeling and simulation, and 3D printing, and parallel cluster computing. She is currently employed as Assistant Professor.
 Mohana Rupa, obtained her Master’s degree in Biotechnology from Oxford College of Science, Bangalore University and Bachelor’s degree from Sree Vidyanikethan Degree College, Sri Venkateswara University. Post her Master’s degree she worked as a research assistant at Sri Padmavathi Mahila Visvavidyalayam, for three years where, her project was focused on understanding the impact of probiotic feed on accumulation and function of antimicrobial peptides in Fish species. As part of her research work, she mastered the process of isolating and purifying proteins from probiotic bacteria using chromatography and other techniques and also in understanding the biological activity of these proteins in enhancing immune responses in Fishes. Currently, she is pursuing her Doctoral degree from VTU – Belagavi, where her work is focused on understanding the structural, functional and genetic characterization of Indian and Western Cattle breeds through comprehensive approach. Her work involves characterizing the structural and functional aspects of milk proteins and their biological significance in normal human health and diseased conditions.
Mohana Rupa, obtained her Master’s degree in Biotechnology from Oxford College of Science, Bangalore University and Bachelor’s degree from Sree Vidyanikethan Degree College, Sri Venkateswara University. Post her Master’s degree she worked as a research assistant at Sri Padmavathi Mahila Visvavidyalayam, for three years where, her project was focused on understanding the impact of probiotic feed on accumulation and function of antimicrobial peptides in Fish species. As part of her research work, she mastered the process of isolating and purifying proteins from probiotic bacteria using chromatography and other techniques and also in understanding the biological activity of these proteins in enhancing immune responses in Fishes. Currently, she is pursuing her Doctoral degree from VTU – Belagavi, where her work is focused on understanding the structural, functional and genetic characterization of Indian and Western Cattle breeds through comprehensive approach. Her work involves characterizing the structural and functional aspects of milk proteins and their biological significance in normal human health and diseased conditions.
Email: rupa.mohana1@gmail.com
